2026-04-28
95% Accuracy That Drops to 58%: Why Deep Learning Models Fail on Biological Signals
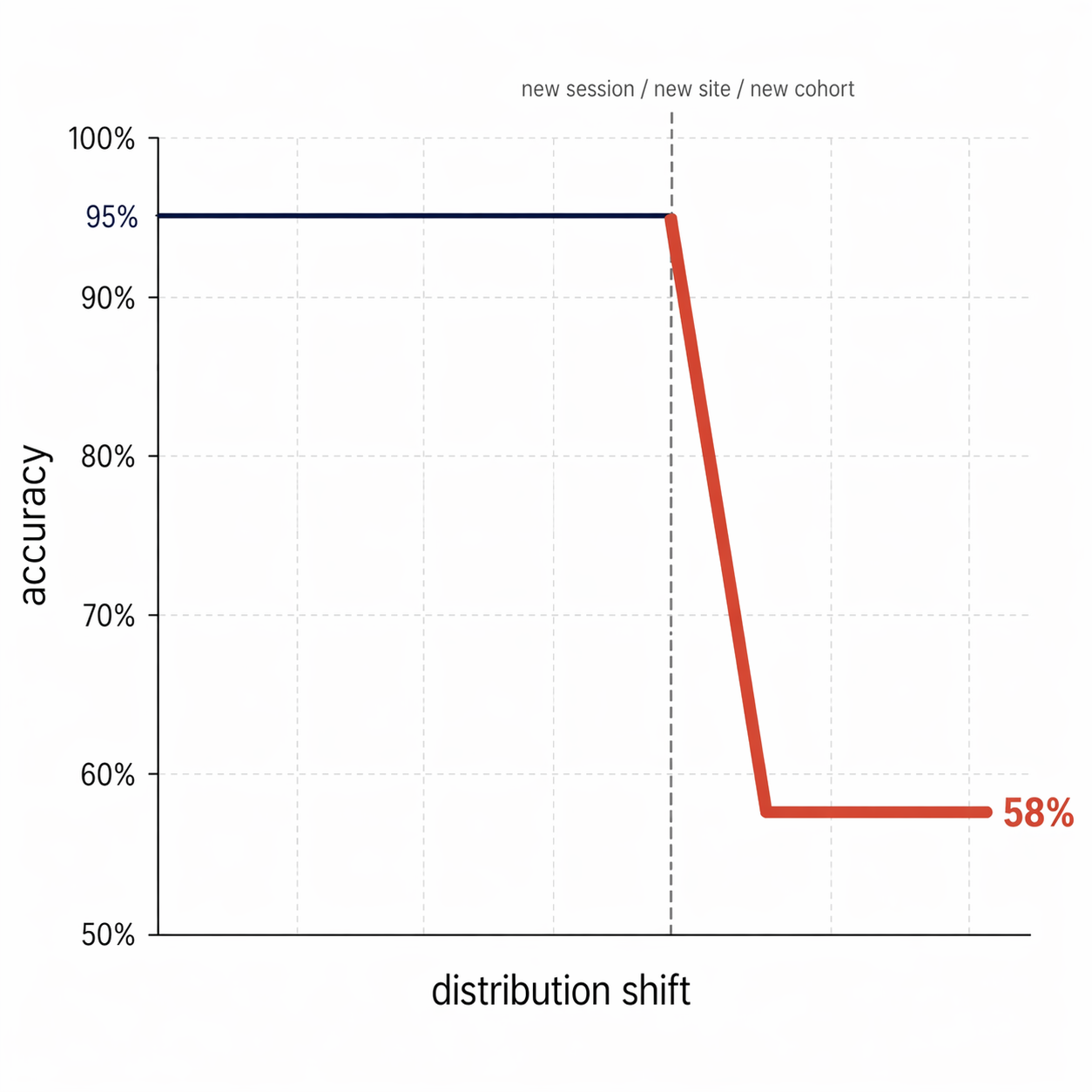
A 95% accuracy model that drops to 58% when the experimental setup changes
I've seen it happen. And it was predictable.
My field was neural signals. But the problem reaches well beyond electrophysiology. It hits every deep learning model applied to biological signals: ECG, EEG, sensor data, dynamic imaging, preclinical data.
The test set lies more often than you think
The test set score lies, because the model isn't learning biology. It's learning the signature of your acquisition session.
As long as the test segments come from the same sessions as the training segments, the model is just recognizing a context it has already seen. The reported performance measures memorization, not understanding.
What shifts when a single variable moves
- Change the device: the noise spectrum shifts
- Change the preparation or the patient: amplitude varies
- Change the site, the center, the cohort: the distribution explodes
- Change the annotator: the labels move
The model never learned to separate signal from context. It mapped a specific context.
The only metric that actually matters
For a model destined for production or a clinical trial, the metric that matters isn't in-distribution accuracy.
It's cross-session, cross-subject, cross-site generalization.
That's what decides whether the model survives the move from lab to real world — from a research protocol to clinical use, from a pilot cohort to a multi-site deployment.
Three shifts that change practice
1. Split by full sessions, not by segments
The train/test boundary has to isolate entire sessions, subjects, or sites. If a single segment from a session ends up in both train and test, the leak is already there — and your measured accuracy is a mirage.
2. Have a domain expert validate the learned features
Grad-CAM, wavelets, saliency maps: the tools exist to open the black box. But the biological reading of what they reveal has to come from a signal expert — not from the data scientist alone. If the model is leaning on a filtering artifact or an amplifier transient, only the domain eye will catch it.
3. Treat the acquisition protocol as part of the architecture
The acquisition protocol matters as much as the convolutional layer that follows it. Controlled session variability, site randomization, annotator balancing: those upstream decisions determine what the model can generalize. A robust model in biology starts with a robust data protocol.
Where is the leak hiding in your pipeline?
If you split by segments, the leak is almost guaranteed. By sessions, it's limited but not eliminated. By subjects, you're truly testing generalization — but the test subjects need to come from different cohorts or sites for the result to translate to scale.
If you have a biological signal ML pipeline to audit — or a model you suspect has learned context rather than signal — take 30 minutes to talk it through.